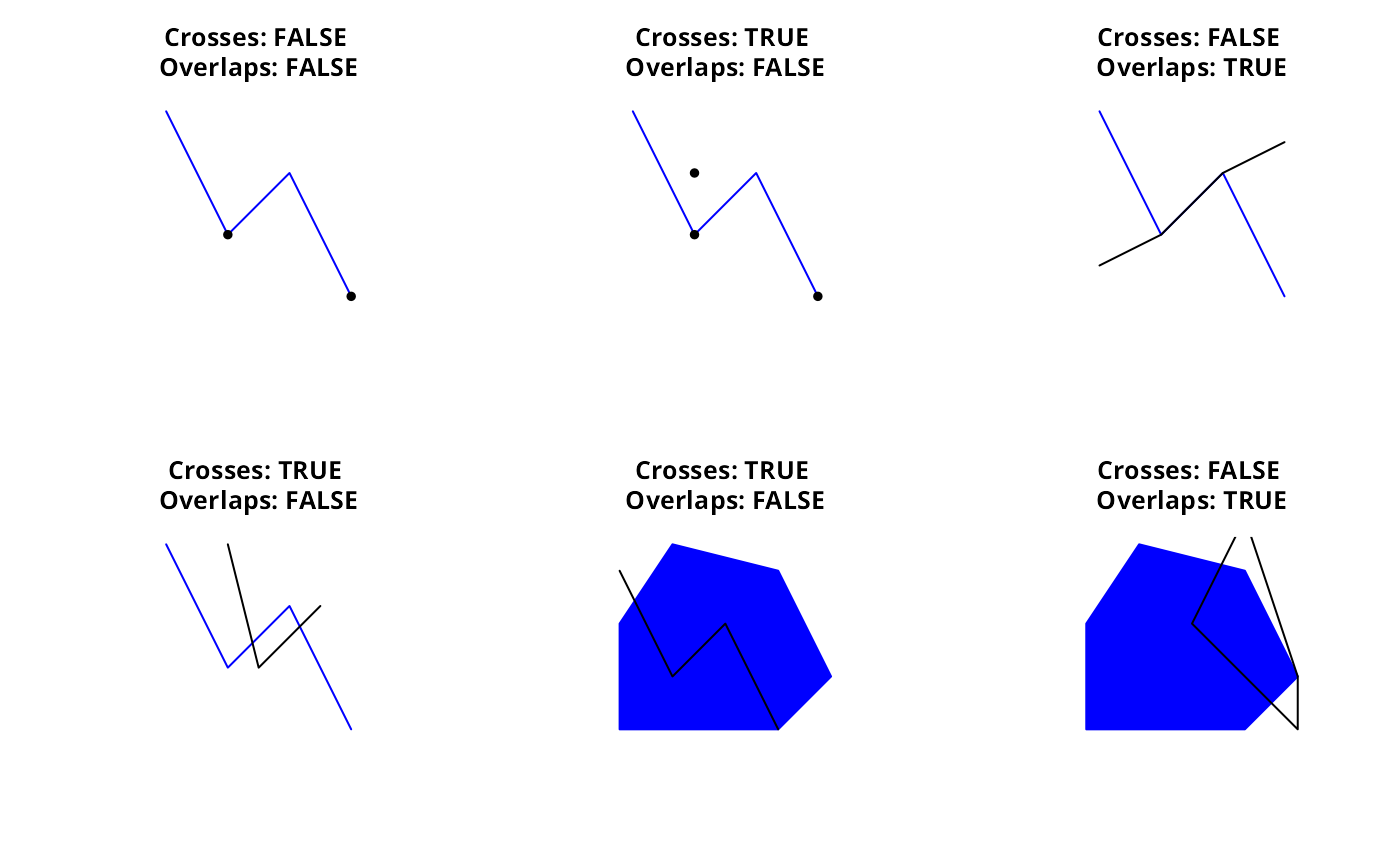
Geometry Relationships - Crosses and Overlaps
pred-binary-gCrosses.RdFunctions for testing whether geometries share some but not all interior points
gCrosses(spgeom1, spgeom2 = NULL, byid = FALSE, returnDense=TRUE,
checkValidity=FALSE)
gOverlaps(spgeom1, spgeom2 = NULL, byid = FALSE, returnDense=TRUE,
checkValidity=FALSE)Arguments
- spgeom1, spgeom2
sp objects as defined in package sp. If spgeom2 is NULL then spgeom1 is compared to itself.
- byid
Logical vector determining if the function should be applied across ids (TRUE) or the entire object (FALSE) for spgeom1 and spgeom2
- returnDense
default TRUE, if false returns a list of the length of spgeom1 of integer vectors listing the
1:length(spgeom2)indices which would be TRUE in the dense logical matrix representation; useful when the sizes of the byid=TRUE returned matrix is very large and it is sparse; essential when the returned matrix would be too large- checkValidity
default FALSE; error meesages from GEOS do not say clearly which object fails if a topology exception is encountered. If this argument is TRUE,
gIsValidis run on each in turn in an environment in which object names are available. If objects are invalid, this is reported and those affected are named
Value
gCrosses returns TRUE when the geometries share some but not all interior points, and the dimension of the intersection is less than that of at least one of the geometries.
gOverlaps returns TRUE when the geometries share some but not all interior points, and the intersection has the same dimension as the geometries themselves.
Note
Error messages from GEOS, in particular topology exceptions, report 0-based object order, so geom 0 is spgeom1, and geom 1 is spgeom2.
See also
Examples
l1 = readWKT("LINESTRING(0 3,1 1,2 2,3 0)")
l2 = readWKT("LINESTRING(0 0.5,1 1,2 2,3 2.5)")
l3 = readWKT("LINESTRING(1 3,1.5 1,2.5 2)")
pt1 = readWKT("MULTIPOINT(1 1,3 0)")
pt2 = readWKT("MULTIPOINT(1 1,3 0,1 2)")
p1 = readWKT("POLYGON((0 0,0 2,1 3.5,3 3,4 1,3 0,0 0))")
p2 = readWKT("POLYGON((2 2,3 4,4 1,4 0,2 2))")
par(mfrow=c(2,3))
plot(l1,col='blue');plot(pt1,add=TRUE,pch=16)
title(paste("Crosses:",gCrosses(l1,pt1),
"\nOverlaps:",gOverlaps(l1,pt1)))
plot(l1,col='blue');plot(pt2,add=TRUE,pch=16)
title(paste("Crosses:",gCrosses(l1,pt2),
"\nOverlaps:",gOverlaps(l1,pt2)))
plot(l1,col='blue');plot(l2,add=TRUE)
title(paste("Crosses:",gCrosses(l1,l2),
"\nOverlaps:",gOverlaps(l1,l2)))
plot(l1,col='blue');plot(l3,add=TRUE)
title(paste("Crosses:",gCrosses(l1,l3),
"\nOverlaps:",gOverlaps(l1,l3)))
plot(p1,border='blue',col='blue');plot(l1,add=TRUE)
title(paste("Crosses:",gCrosses(p1,l1),
"\nOverlaps:",gOverlaps(p1,l1)))
plot(p1,border='blue',col='blue');plot(p2,add=TRUE)
title(paste("Crosses:",gCrosses(p1,p2),
"\nOverlaps:",gOverlaps(p1,p2)))